吴裕雄--天生自然 PYTHON数据分析:糖尿病视网膜病变数据分析(续四)
内容导读
互联网集市收集整理的这篇技术教程文章主要介绍了吴裕雄--天生自然 PYTHON数据分析:糖尿病视网膜病变数据分析(续四),小编现在分享给大家,供广大互联网技能从业者学习和参考。文章包含4206字,纯文字阅读大概需要7分钟。
内容图文
dpi = 80 #inch
path_jpg=f"F:\\kaggleDataSet\\diabeticRetinopathy\\resized_train_cropped\\18017_left.jpeg" # too many vessels?
path_png=f"F:\\kaggleDataSet\\diabeticRetinopathy\\rescaled_train_896\\18017_left.png" # details are lost
image = cv2.imread(path_png)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image = cv2.resize(image, (IMG_SIZE, IMG_SIZE))
image2 = cv2.imread(path_jpg)
image2 = cv2.cvtColor(image2, cv2.COLOR_BGR2RGB)
image2 = cv2.resize(image2, (IMG_SIZE, IMG_SIZE))
height, width = IMG_SIZE, IMG_SIZE
print(height, width)
SCALE=1/4
figsize = (width / float(dpi))/SCALE, (height / float(dpi))/SCALE
fig = plt.figure(figsize=figsize)
ax = fig.add_subplot(2, 2, 1, xticks=[], yticks=[])
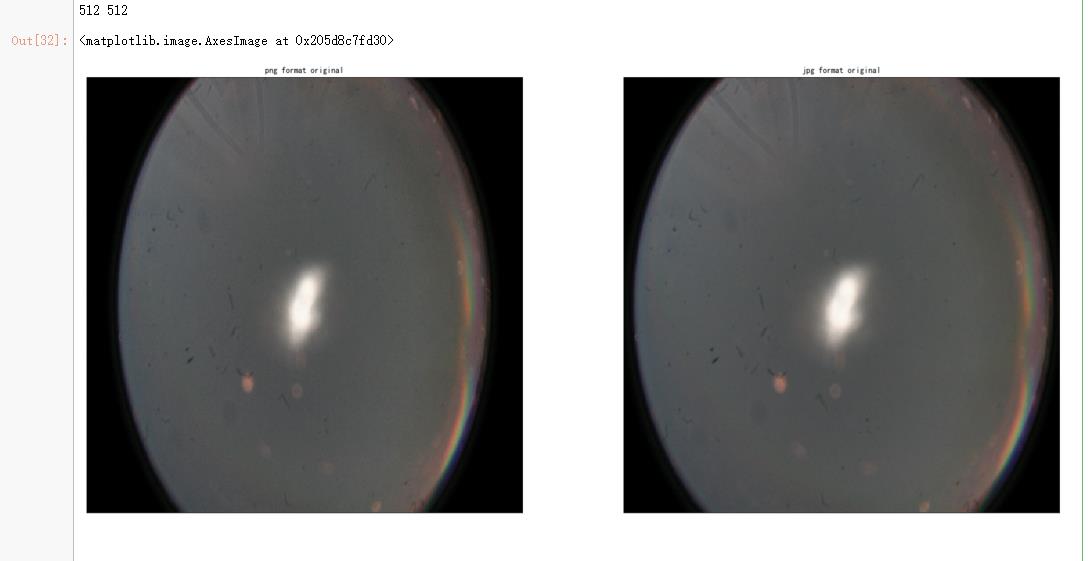
ax.set_title('png format original' )
plt.imshow(image, cmap='gray')
ax = fig.add_subplot(2, 2, 2, xticks=[], yticks=[])
ax.set_title('jpg format original' )
plt.imshow(image2, cmap='gray')
image = load_ben_color(path_png,sigmaX=30)
image2 = load_ben_color(path_jpg,sigmaX=30)
ax = fig.add_subplot(2, 2, 3, xticks=[], yticks=[])
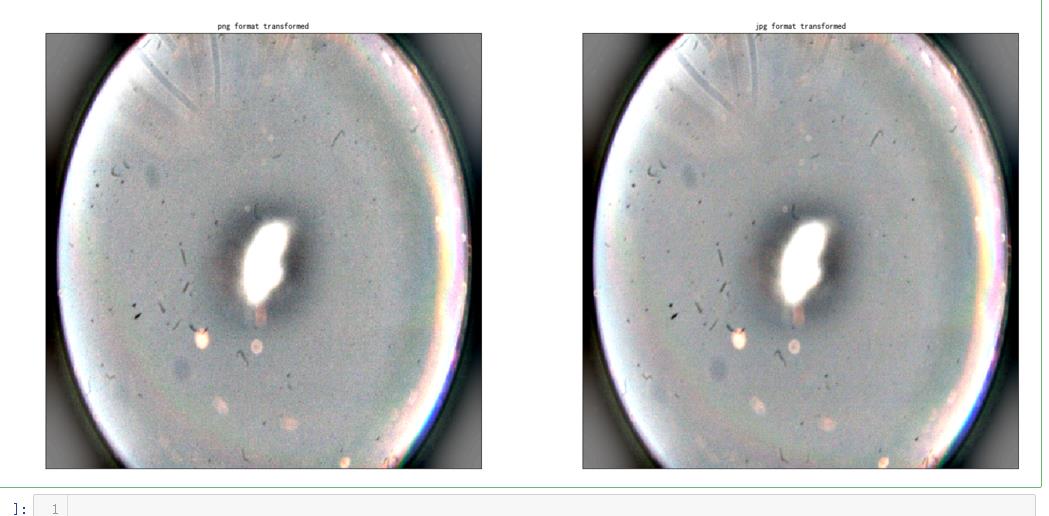
ax.set_title('png format transformed' )
plt.imshow(image, cmap='gray')
ax = fig.add_subplot(2, 2, 4, xticks=[], yticks=[])
ax.set_title('jpg format transformed' )
plt.imshow(image2, cmap='gray')
import json import math import os import cv2 from PIL import Image import numpy as np from keras import layers from keras.applications import DenseNet121 from keras.callbacks import Callback, ModelCheckpoint from keras.preprocessing.image import ImageDataGenerator from keras.models import Sequential from keras.optimizers import Adam import matplotlib.pyplot as plt import pandas as pd from sklearn.model_selection import train_test_split from sklearn.metrics import cohen_kappa_score, accuracy_score import scipy from tqdm import tqdm %matplotlib inline
train_df = pd.read_csv('F:\\kaggleDataSet\\diabeticRetinopathy\\trainLabels19.csv')
test_df = pd.read_csv('F:\\kaggleDataSet\\diabeticRetinopathy\\testImages19.csv')
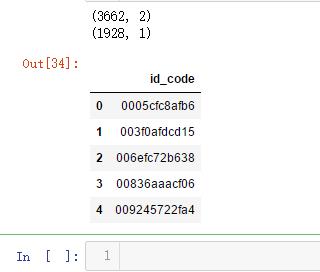
print(train_df.shape)
print(test_df.shape)
test_df.head()
def get_pad_width(im, new_shape, is_rgb=True):
pad_diff = new_shape - im.shape[0], new_shape - im.shape[1]
t, b = math.floor(pad_diff[0]/2), math.ceil(pad_diff[0]/2)
l, r = math.floor(pad_diff[1]/2), math.ceil(pad_diff[1]/2)
if is_rgb:
pad_width = ((t,b), (l,r), (0, 0))
else:
pad_width = ((t,b), (l,r))
return pad_width
def preprocess_image(image_path, desired_size=224):
im = Image.open(image_path)
im = im.resize((desired_size, )*2, resample=Image.LANCZOS)
return im
N = test_df.shape[0]
x_test = np.empty((N, 224, 224, 3), dtype=np.uint8)
for i, image_id in enumerate(tqdm(test_df['id_code'])):
x_test[i, :, :, :] = preprocess_image("F:\\kaggleDataSet\\diabeticRetinopathy\\resized test 19\\"+str(image_id)+".jpg")
# model.summary()
def load_image_ben_orig(path,resize=True,crop=False,norm255=True,keras=False):
image = cv2.imread(path)
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
image=cv2.addWeighted( image,4, cv2.GaussianBlur( image , (0,0) , 10) ,-4 ,128)
if norm255:
return image/255
elif keras:
#see https://github.com/keras-team/keras-applications/blob/master/keras_applications/imagenet_utils.py for mode
#see https://github.com/keras-team/keras-applications/blob/master/keras_applications/xception.py for inception,xception mode
#the use of tf based preprocessing (- and / by 127 respectively) will results in [-1,1] so it will not visualize correctly (directly)
image = np.expand_dims(image, axis=0)
return preprocess_input(image)[0]
else:
return image.astype(np.int16)
return image
def transform_image_ben(img,resize=True,crop=False,norm255=True,keras=False):
image=cv2.addWeighted( img,4, cv2.GaussianBlur( img , (0,0) , 10) ,-4 ,128)
if norm255:
return image/255
elif keras:
image = np.expand_dims(image, axis=0)
return preprocess_input(image)[0]
else:
return image.astype(np.int16)
return image
def display_samples(df, columns=5, rows=2, Ben=True):
fig=plt.figure(figsize=(5*columns, 4*rows))
for i in range(columns*rows):
image_path = df.loc[i,'id_code']
path = f"F:\\kaggleDataSet\\diabeticRetinopathy\\resized test 19\\"+str(image_path)+".jpg"
if Ben:
img = load_image_ben_orig(path)
else:
img = cv2.imread(path)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
fig.add_subplot(rows, columns, i+1)
plt.imshow(img)
plt.tight_layout()
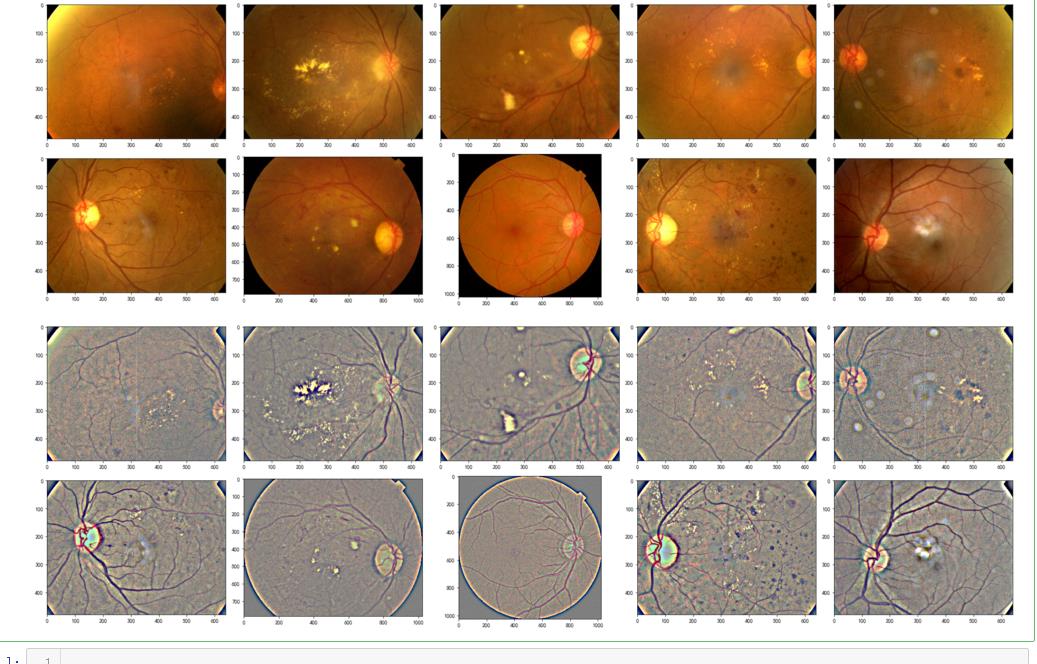
display_samples(test_df, Ben=False)
display_samples(test_df, Ben=True)
内容总结
以上是互联网集市为您收集整理的吴裕雄--天生自然 PYTHON数据分析:糖尿病视网膜病变数据分析(续四)全部内容,希望文章能够帮你解决吴裕雄--天生自然 PYTHON数据分析:糖尿病视网膜病变数据分析(续四)所遇到的程序开发问题。 如果觉得互联网集市技术教程内容还不错,欢迎将互联网集市网站推荐给程序员好友。
内容备注
版权声明:本文内容由互联网用户自发贡献,该文观点与技术仅代表作者本人。本站仅提供信息存储空间服务,不拥有所有权,不承担相关法律责任。如发现本站有涉嫌侵权/违法违规的内容, 请发送邮件至 gblab@vip.qq.com 举报,一经查实,本站将立刻删除。
内容手机端
扫描二维码推送至手机访问。